doi: 10.56294/dm2024.353
SHORT COMMUNICATION
Variability of the SNP rs9939609 in the FTO Gene and Ancestry in Latin American Populations
Variabilidad del SNP rs9939609 en el Gen FTO y la Ancestría en Poblaciones Latinoamericanas
Sergio V. Flores1
![]() *, Román M.
Montaña2
*, Román M.
Montaña2 ![]() *, Angel
Roco-Videla3
*, Angel
Roco-Videla3 ![]() *, Marcela
Caviedes-Olmos4
*, Marcela
Caviedes-Olmos4 ![]() *, Raúl Aguilera
Eguía5
*, Raúl Aguilera
Eguía5 ![]() *
*
1Universidad Arturo Prat. Santiago, Chile.
2Universidad Autónoma de Chile, Facultad de Ciencias de la Salud. Santiago, Chile.
3Universidad Bernardo O´Higgins, Programa de Magister en ciencias químico-Biológicas. Santiago, Chile.
4Universidad de las Américas. Facultad de Salud y Ciencias Sociales, Santiago, Chile.
5Universidad Católica de la Santísima Concepción, Departamento de Salud Pública, Concepción, Chile.
Cite as: Flores SV, Montaña RM, Roco-Videla A, Caviedes-Olmos M, Aguilera Eguía R. Variability of the SNP rs9939609 in the FTO Gene and Ancestry in Latin American Populations. Data and Metadata. 2024;3:.353. https://doi.org/10.56294/dm2024.353
Submitted: 19-01-2024 Revised: 03-05-2024 Accepted: 20-09-2024 Published: 21-09-2024
Editor: Adrián
Alejandro Vitón Castillo
![]()
Corresponding author: Sergio V. Flores1 *
ABSTRACT
Introduction: obesity is a complex condition influenced by genetic and environmental factors. The FTO gene has been associated with obesity through several single nucleotide polymorphisms (SNPs), particularly rs9939609, related to higher body mass index (BMI) and risk of obesity. FTO variants influence the regulation of appetite and energy metabolism by affecting RNA methylation and the expression of key genes in adipogenesis.
Objective: to investigate the association between the FTO rs9939609 SNP and genetic ancestry proportions in Latin American populations.
Methods: genotypes for rs9939609 were obtained using VcfTools and the 1000 Genomes Project database. Samples from Latin America were selected, covering four mixed populations: Colombians (n=94), Mexicans (n=64), Peruvians (n=85) and Puerto Ricans (n=104), totaling 347 individuals. To estimate genetic ancestry proportions, 446 SNPs from a panel of ancestry informative markers (AIMs) were used.
Results: individuals with the AA genotype of SNP rs9939609 have a higher proportion of Native American ancestry and a lower proportion of European ancestry compared to TT and AT genotypes. The variability in the proportions of ancestry according to the genotype of the SNP rs9939609 suggests a possible genetic stratification in the Latin American populations studied.
Conclusions: these findings highlight the importance of considering ancestral composition in genetic studies related to obesity. More research is needed to understand how gene-environment interactions contribute to obesity in various populations, which could lead to more effective and targeted intervention strategies.
Keywords: Alpha-Ketoglutarate-Dependent Dioxygenase FTO; Obesity; Latin America; Body Mass Index.
RESUMEN
Introducción: la obesidad es una condición compleja influenciada por factores genéticos y ambientales. El gen FTO se ha asociado con la obesidad a través de varios polimorfismos de nucleótido único (SNPs), particularmente rs9939609, relacionado con un mayor índice de masa corporal (IMC) y riesgo de obesidad. Las variantes de FTO influyen en la regulación del apetito y el metabolismo energético al afectar la metilación del ARN y la expresión de genes clave en la adipogénesis.
Objetivo: investigar la asociación entre el SNP rs9939609 del gen FTO y las proporciones de ascendencia genética en poblaciones latinoamericanas.
Métodos: se obtuvieron los genotipos para rs9939609 utilizando VcfTools y la base de datos del Proyecto 1000 Genomas. Se seleccionaron muestras de América Latina, abarcando cuatro poblaciones mezcladas: colombianos (n=94), mexicanos (n=64), peruanos (n=85) y puertorriqueños (n=104), totalizando 347 individuos. Para estimar las proporciones de ancestría genética, se utilizaron 446 SNPs de un panel de marcadores informativos de ancestría (AIMs).
Resultados: los individuos con el genotipo AA del SNP rs9939609 tienen una mayor proporción de ancestría nativo americana y una menor proporción de ancestría europea en comparación con los genotipos TT y AT. La variabilidad en las proporciones de ancestría según el genotipo del SNP rs9939609 sugiere una posible estratificación genética en las poblaciones latinoamericanas estudiadas.
Conclusiones: estos hallazgos resaltan la importancia de considerar la composición ancestral en los estudios genéticos relacionados con la obesidad. Se necesita más investigación para comprender cómo la interacción entre genes y ambiente contribuye a la obesidad en varias poblaciones, lo que podría conducir a estrategias de intervención más efectivas y específicas.
Palabras clave: Dioxigenasa FTO Dependiente De Alfa-Cetoglutarato; Rs9939609; Obesidad; Ancestría Genética; América Latina; Índice De Masa Corporal.
INTRODUCTION
The FTO gene (fat mass and obesity-associated) is associated with obesity through various single nucleotide polymorphisms (SNPs), particularly rs9939609, which is related to higher body mass index (BMI) and obesity risk.(1) Variants of FTO influence appetite regulation and energy metabolism by affecting RNA methylation and the expression of key genes in adipogenesis.(2, 3) Individuals with risk alleles tend to consume more energy-dense foods, leading to increased caloric intake and weight gain.(4)
The FTO gene is also associated with cancer through its role in the demethylation of N6-methyladenosine (m6A) in various types of RNA.(5) SNPs in FTO, such as rs9939609, are linked to higher BMI and predisposition to obesity.(1, 4) These SNPs influence the expression of key genes in adipogenesis and appetite control, such as RUNX1T1 and ghrelin.(6,7) Additionally, FTO regulates the mTOR pathway, which is important for cell growth and metabolism, and its dysfunction can contribute to cancer development.(8,9) FTO is overexpressed in various types of cancer, such as leukemia, glioblastoma, and breast cancer, where its m6A demethylase activity influences tumor progression and treatment resistance, suggesting that FTO acts as a molecular link between obesity and cancer.(3,10,11)
The FTO gene shows significant population variability, especially in the SNP rs9939609, which has shown a strong correlation with BMI in individuals of European and Asian ancestry.(1,4) The risk allele A of rs9939609 is associated with higher BMI and obesity risk in Europe and Asia.(12,13) These associations are also observed in populations of Latin America and Africa.(14,15)
The aim of this research is to analyze the relationship between European ancestry and the prevalence of the risk allele rs9939609 in Latin American populations, to better understand how genetic variability influences obesity risk. The Latin American population, descended from East Asian and European populations, presents a complex genetic structure. The rate of recombination and genetic structuring, influenced by sociocultural segregation, could be causing stratification of FTO alleles according to the degree of European ancestry.
METHODS
Genotypes for rs9939609 were obtained using VcfTools(16) and the 1000 Genomes Project Consortium database. (17) For this study, samples exclusively from Latin America were selected, encompassing four admixed populations: Colombians (n=94), Mexicans (n=64), Peruvians (n=85), and Puerto Ricans (n=104), totaling 347 individuals.
To estimate genetic ancestry proportions, 446 SNPs from a panel of ancestry informative markers (AIMs) recommended by Galanter et al. were used.(18) These markers were designed and validated to calculate ancestry proportions in individuals and populations in Latin America.
Five ancestral populations (K=5) were modeled to estimate the genetic ancestry proportions of each individual using STRUCTURE.(19) The Kolmogorov-Smirnov test was then applied to assess the normality of the distribution of individual genetic ancestry proportions. Since none of the ancestries followed a normal distribution (P Kolmogorov-Smirnov < 0,05), Non-parametric tests were used: Kruskal-Wallis and post hoc Wilcoxon pairwise tests, to evaluate the null hypothesis of no association between genetic ancestry and the obesity risk genotypes, with a confidence level of 95 % and a critical p-value of 0,05.
RESULTS
The distribution and normality analysis of Native American (NAM) and European (EUR) ancestries were performed using histograms, Q-Q plots (figure 1), and Shapiro-Wilk normality tests. The NAM ancestry distribution showed significant skewness, confirmed by a Shapiro-Wilk statistic of 0,8787 and a p-value of 6,63×10−16. The EUR ancestry showed a distribution closer to normality, with a Shapiro-Wilk statistic of 0,9653 and a p-value of 2,40×10−7.
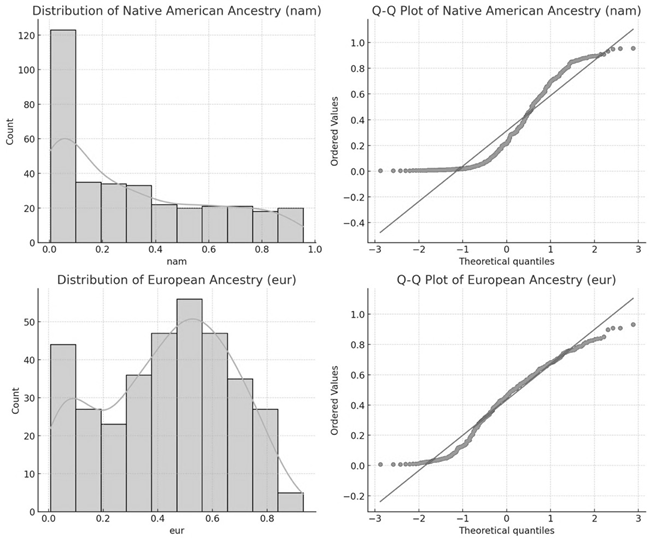
Figure 1. Distribution of Native-American ancestry and European ancestry proportions among individuals
Kruskal-Wallis tests revealed significant differences in NAM and EUR ancestry proportions among the TT, AT, and AA genotypes of the SNP rs9939609. For NAM, the Kruskal-Wallis statistic was 48,913 (p = 2,39×10−11), while for EUR, it was 38,820 (p = 3,72×10−9). Box plots showed that the AA genotype has a higher median and a wider range in NAM, and a lower median in EUR, compared to the TT and AT genotypes (figure 2).
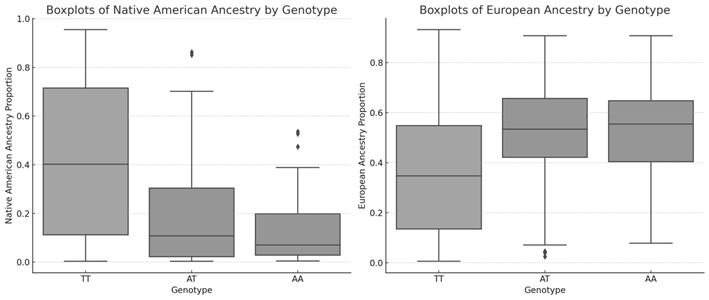
Figure 2. Distribution of European ancestry (left) and Native-American ancestry (right) among the genotypes of the rs4988235(C) polymorphism
Post-hoc Mann-Whitney U tests showed significant differences in NAM ancestry between TT and AT genotypes (U = 17866,5, p = 9,03×10−11), and between TT and AA genotypes (U = 3537,5, p = 7,07×10−5). For EUR ancestry, there were significant differences between TT and AT genotypes (U = 7699,0, p = 3,83×10−9) and between TT and AA genotypes (U = 1422,5, p = 0,00113). No significant differences were found between AT and AA genotypes in any ancestry.
DISCUSSION
The A allele of the SNP rs9939609 in the FTO gene is associated with a higher likelihood of obesity, a complex condition influenced by both genetic and environmental factors.(1) The results of this study show significant differences in Native American and European ancestry proportions among the different genotypes of this SNP. Individuals with the AA genotype (low risk of obesity) have a higher proportion of Native American ancestry and a lower proportion of European ancestry compared to those with TT and AT genotypes. This suggests that variation in the FTO gene and its association with obesity may differ among populations with different ancestral compositions.
The higher frequency of the A allele in European populations could partly explain the differences in obesity prevalence between these and other populations.(12) Individuals with the A allele have a genetic predisposition that affects appetite regulation and metabolism, increasing the risk of developing obesity.(2,3) The influence of genotype on the variability of ancestral composition observed in this study highlights the importance of considering specific genetic factors in each population when studying complex diseases like obesity.
These findings underscore the need for personalized approaches in the treatment and prevention of obesity, considering both individual genetics and ancestral diversity.(4,5) Future research should focus on how the interaction between genes and the environment contributes to obesity in various populations, which could lead to more effective and targeted intervention strategies.
Finally, some limitations of this study are related to the restriction to a sample of four Latin American populations, which may not represent the complete genetic diversity of the region. The observed genetic stratification may be affected by sample selection and the accuracy of ancestry estimates. Nonetheless, the alignment of the results with predictions suggests expanding the study’s approach to other populations.
CONCLUSIONS
Individuals with the AA genotype of the SNP rs9939609 have a higher proportion of Native American ancestry and a lower proportion of European ancestry compared to the TT and AT genotypes. The variability in ancestry proportions according to the genotype of the SNP rs9939609 suggests potential genetic stratification in the studied Latin American populations. These findings highlight the importance of considering ancestral composition in genetic studies related to obesity.
REFERENCES
1. Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 2007;316:889–94. https://doi.org/10.1126/science.1141634
2. Cecil JE, Tavendale R, Watt P, Hetherington MM, Palmer CNA. An obesity-associatedFTOgene variant and increased energy intake in children. N Engl J Med 2008;359:2558–66. https://doi.org/10.1056/nejmoa0803839
3. Lan N, Lu Y, Zhang Y, Pu S, Xi H, Nie X, et al. FTO – A common genetic basis for obesity and cancer. Front Genet 2020;11. https://doi.org/10.3389/fgene.2020.559138
4. Loos RJF, Yeo GSH. The bigger picture of FTO—the first GWAS-identified obesity gene. Nat Rev Endocrinol 2014;10:51–61. https://doi.org/10.1038/nrendo.2013.227
5. Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, et al. N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 2011;7:885–7. https://doi.org/10.1038/nchembio.687
6. Karra E, O’Daly OG, Choudhury AI, Yousseif A, Millership S, Neary MT, et al. A link between FTO, ghrelin, and impaired brain food-cue responsivity. J Clin Invest 2013;123:3539–51. https://doi.org/10.1172/jci44403
7. Zhao X, Yang Y, Sun B-F, Shi Y, Yang X, Xiao W, et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res 2014;24:1403–19. https://doi.org/10.1038/cr.2014.151
8. Cheung MK, Gulati P, O’Rahilly S, Yeo GSH. FTO expression is regulated by availability of essential amino acids. Int J Obes (Lond) 2013;37:744–7. https://doi.org/10.1038/ijo.2012.77.
9. Gulati P, Cheung MK, Antrobus R, Church CD, Harding HP, Tung Y-CL, et al. Role for the obesity-related FTO gene in the cellular sensing of amino acids. Proc Natl Acad Sci U S A 2013;110:2557–62. https://doi.org/10.1073/pnas.1222796110.
10. Niu Y, Lin Z, Wan A, Chen H, Liang H, Sun L, et al. RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Mol Cancer 2019;18. https://doi.org/10.1186/s12943-019-1004-4.
11. Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, et al. M 6 A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep 2017;18:2622–34. https://doi.org/10.1016/j.celrep.2017.02.059.
12. Dina C, Meyre D, Gallina S, Durand E, Körner A, Jacobson P, et al. Variation in FTO contributes to childhood obesity and severe adult obesity. Nat Genet 2007;39:724–6. https://doi.org/10.1038/ng2048.
13. Wen W, The Genetic Investigation of ANthropometric Traits (GIANT) Consortium, Cho Y-S, Zheng W, Dorajoo R, Kato N, et al. Meta-analysis identifies common variants associated with body mass index in east Asians. Nat Genet 2012;44:307–11. https://doi.org/10.1038/ng.1087.
14. da Fonseca ACP, Abreu GM, Zembrzuski VM, Campos Junior M, Carneiro JRI, Nogueira Neto JF, et al. The association of the fat mass and obesity-associated gene (FTO) rs9939609 polymorphism and the severe obesity in a Brazilian population. Diabetes Metab Syndr Obes 2019;12:667–84. https://doi.org/10.2147/dmso.s199542.
15. Jiang Y, Qi X, Liu X, Zhang J, Liu T. Association of FTO rs9939609 polymorphism with obesity risk in children: A meta-analysis. World J Pediatr. 2019;15(3):281-91.
16. Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, et al. The variant call format and VCFtools. Bioinformatics 2011;27:2156–8. https://doi.org/10.1093/bioinformatics/btr330.
17. The 1000 Genomes Project Consortium, Auton A, Abecasis GR, Altshuler DM (Co-Chair), Durbin RM (Co-Chair), Abecasis GR, et al. A global reference for human genetic variation. Nature 2015;526:68–74. https://doi.org/10.1038/nature15393.
18. Galanter JM, Fernandez-Lopez JC, Gignoux CR, Barnholtz-Sloan J, Fernandez-Rozadilla C, Via M, et al. Development of a panel of genome-wide ancestry informative markers to study admixture throughout the Americas. PLoS Genet 2012;8:e1002554. https://doi.org/10.1371/journal.pgen.1002554.
19. Hubisz MJ, Falush D, Stephens M, Pritchard JK. Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 2009;9:1322–32. https://doi.org/10.1111/j.1755-0998.2009.02591.x.
FINANCING
The authors did not receive financing for the development of this research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORSHIP CONTRIBUTION
Conceptualization: Sergio V. Flores
Data curation: Sergio V. Flores, Román Montaña
Formal analysis: Sergio V. Flores, Angel Roco-Videla, Marcela Caviedes-Olmos
Research: Sergio V. Flores, Angel Roco, Román Montaña
Methodology: Sergio V. Flores, Angel Roco-Videla
Software: Sergio V. Flores
Supervision: Sergio V. Flores
Validation: Angel Roco-Videla, Marcela Caviedes-Olmos, Raúl Aguilera Eguía
Display: Román Montaña
Drafting - original draft: Sergio V. Flores
Writing - proofreading and editing: Angel Roco-Videla, Raúl Aguilera Eguía